If you get this, it works, and SearchGUI should run without problems. This should result in output describing what the script does. To test your installation run the search engine executable on the command line. Search Engine Issues - Important: If you have problems with the search engines, please verify that the search engines are working outside of SearchGUI first. This only has to be done once for each SearchGUI version. 
To escape this warning control-click on the file icon and then select “Open.” This will give you the option of opening it regardless of its unidentified source. Unidentified Developer - If you run SearchGUI on a Mac you can get the warning “SearchGUI” can’t be opened because it is from an unidentified developer. (Note that on Linux SearchGUI has to be run from a path not containing spaces). [, %, æ, ø, å, etc? If so, move the whole folder to a different location or rename the folder(s) causing the problem and try again. Unzip the file and try again.ĭoes Not Start III - Is SearchGUI installed in a path containing special characters, i.e. If you get the message “A Java Exception has occurred”, you are most likely trying to run SearchGUI from within the zip file. (You only need the JRE version (and not the JDK version) to run SearchGUI.)ĭoes Not Start II - Have you unzipped the zip file? You need to unzip the file before double clicking the jar file. PeptideShaker is a search engine independent platform for visualization of peptide and protein identification results from multiple search engines.ĭoes Not Start I - Do you have Java installed? Download the latest version of Java here and try again. To visualize and analyze the SearchGUI results we recommend the use of PeptideShaker. From the command line you have to run the msconvert command line separatelty. Note that this option is only available via the graphical user interface. In addition, ThermoRawFileParser is included, which supports out-of-the-box conversion of Thermo raw files into mzML and mgf.įurthermore, by referencing the location of your ProteoWizard installtion you may also provide additional raw file types as input, which will then be converted using msconvert. SearchGUI supports mzML and mgf files as the direct input format for the spectrum files. This functionality is available by default in PeptideShaker. Using such a modification will result in nonsense matches which can be filtered out afterwards. For example, X! Tandem is not compatible with modifications at termini on specific amino acids. Not all modifications are correctly handled by the search engines. Modifications will be available in other instances of SearchGUI and PeptideShaker for the same user/computer. 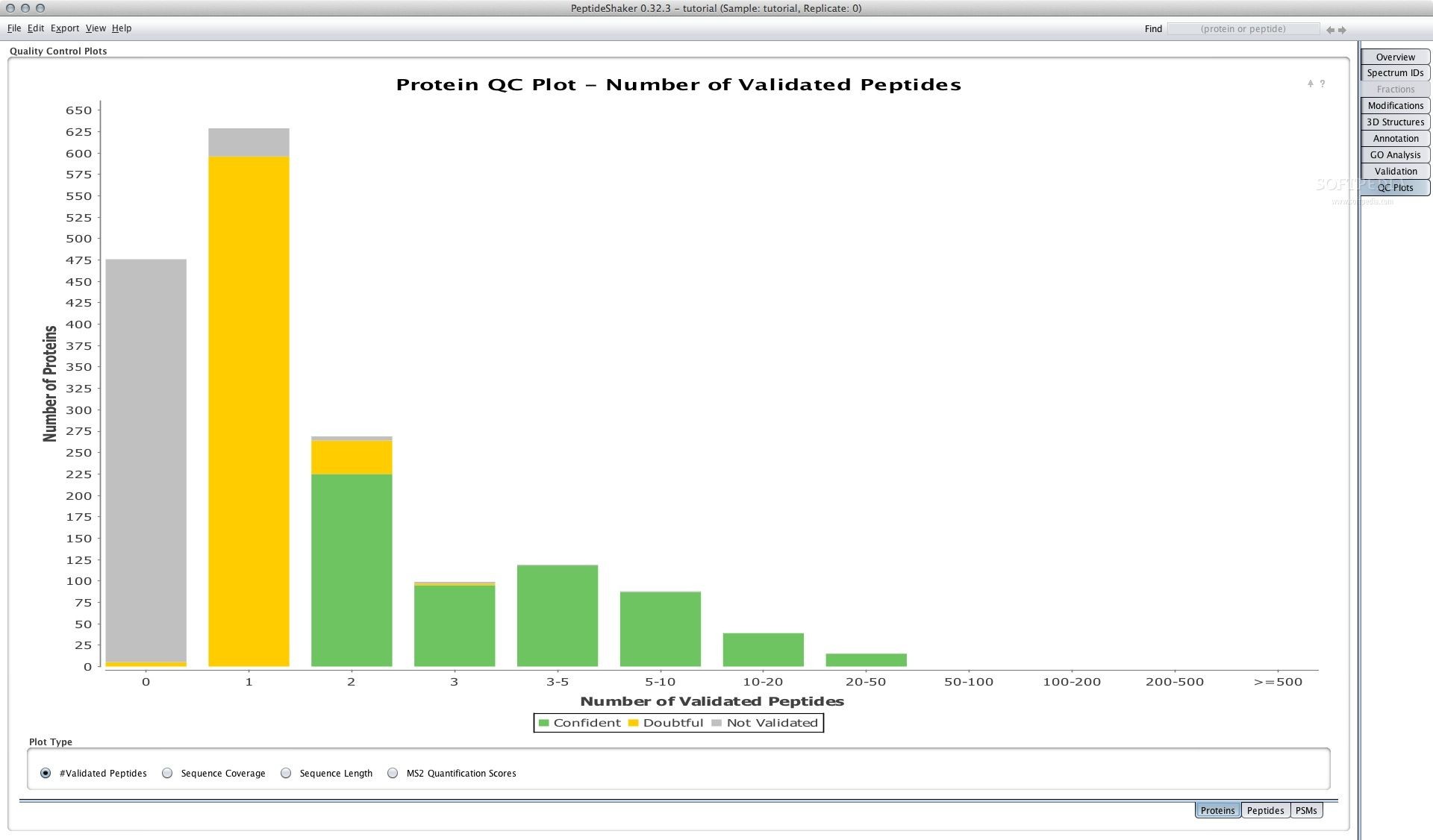
It is straightforward to add/edit modifications via the graphical user interface. SearchGUI is available as a Miniconda package in the bioconda channel here. SearchGUI can also be used via the command line, and be incorporated in different analysis pipelines.įor details about the command line see: SearchCLI. A graphical user interface is the best choice for smaller projects. The main purpose of SearchGUI is to make it simpler to use multiple search engines at the same time. To start identifying peptides and proteins using SearchGUI, download the latest version, unzip the downloaded file, and double-click on the SearchGUI-X.Y.Z.jar file. To visualize and analyze the search results we recommend PeptideShaker.įor developer access to the search results we recommend the use of compomics-utilities. To start using SearchGUI, unzip the downloaded file, and double-click the SearchGUI-X.Y.Z.jar file. SearchGUI is a a highly adaptable open-source common interface for configuring and running proteomics search and de novo engines, currently supporting X! Tandem, MyriMatch, MS Amanda, MS-GF+, OMSSA, Comet, Tide, Andromeda, MetaMorpheus, Sage, Novor and DirecTag. (Click on figure to see the full size version) If you use SearchGUI as part of a publication, please refer to the most recent publication.Vaudel M, Barsnes H, Berven FS, Sickmann A, Martens L: SearchGUI: An open-source graphical user interface for simultaneous OMSSA and X!Tandem searches.Barsnes H and Vaudel M: SearchGUI: a highly adaptable common interface for proteomics search and de novo engines.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
